 
At the end of each chapter, there are exercises and programming challenges offered to reinforce concepts. The author succeeds at providing an informal introduction to Perl scripting that incorporates real-world examples appropriate for the intended audience. PERL PROGRAMMING LANGUAGE HOW TOAdvanced sections on references and object-oriented programming provide the necessary background for understanding how to use Bioperl modules, which are the focus of the last chapter of the book. Intended for biologists, particularly those involved in genomics and proteomics, who have no prior programming experience, this book begins with the basics of data and control structures before covering more intermediate topics of regular expressions and input/output handling. 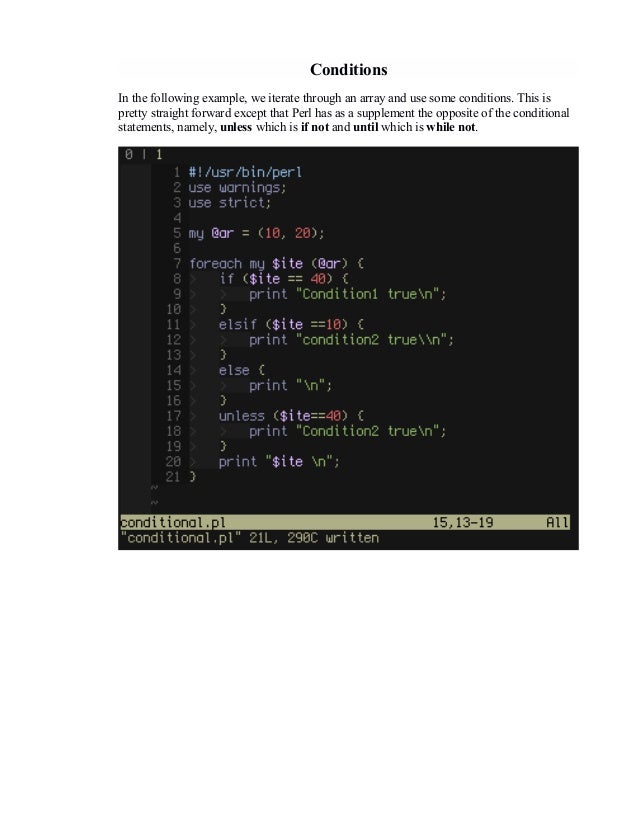
Furthermore, most beginning programmers will find that as their programming projects grow more complex, they will soon need a more thorough treatment of the subject, such as that found in O'Reilly's Beginning Perl for Bioinformatics. This book clearly targets that audience, and in so doing is too elementary for most researchers with intermediate or advanced programming experience. That desire translates into a very accessible text, which any biomedical researcher lacking programming experience would find a quick and painless primer. PERL PROGRAMMING LANGUAGE FULLJamison's long-held desire to see full integration of computer technology into the experimental protocol that he brings to Perl Programming for Biologists. 
The goal of this book is to provide the reader with the background and tools necessary to write scripts of immediate practical value to his or her work. Due in large part to its powerful regular expressions for pattern matching and its accessibility for nonprogrammers, Perl has become the most popular programming language in bioinformatics. Perl is an open-source interpreted scripting language originally designed for Unix systems programming by Larry Wall about 25 years ago. Curtis Jamison, PhD, an associate professor in the School of Computational Science at George Mason University in Manassas, Virginia, wrote the book Perl Programming for Biologists. Originally contributed by Korjavin Ivan, and updated by 10 contributor(s).D. Got a suggestion? A correction, perhaps? Open an Issue on the Github Repo, or make a pull request yourself! Perlfaq contains questions and answers related to many common tasks, and often provides suggestions for good CPAN modules to use. # Scalars # A scalar represents a single value: my $animal = "camel" my $answer = 42 my $display = "You have $answer $ 1 # Object-oriented programming is covered more thoroughly in perlootut, # and its low-level implementation in Perl is covered in perlobj. # Perl has three main variable types: $scalar, and %hash. # A valid variable name starts with a letter or underscore, # followed by any number of letters, numbers, or underscores. 
# Perl variable types # Variables begin with a sigil, which is a symbol showing the type. Strict causes # compilation to fail in cases like misspelled variable names, and # warnings will print warning messages in case of common pitfalls like # concatenating to an undefined value. # Strict and warnings use strict use warnings # All perl scripts and modules should include these lines. # Single line comments start with a number sign. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
